Digital Intelligence for Kidney Diseases and Organ Transplantation
Collaborators: Karthik Tennankore, Amanda Vinson, George Worthen
Funding: NS Health
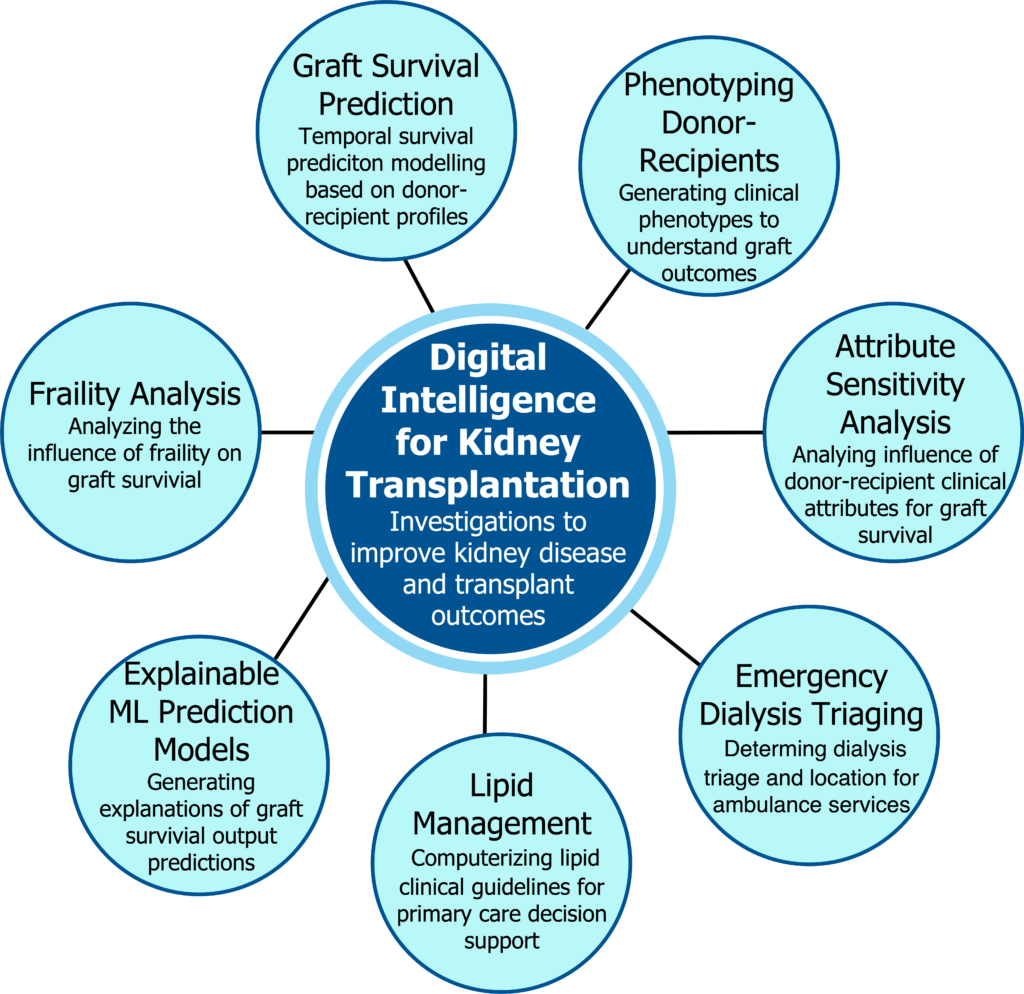
Synopsis: This research program is investigating AI methods for (a) optimizing kidney transplant survival by studying donor and recipients characteristics, and (b) improving treatment of chronic kidney diseases. We are conducting multiple projects as follows:
1. Kidney Transplant Survival Prediction: Kidney transplantation is the optimal form of renal replacement therapy, but patients are still at risk of death or premature kidney graft loss. Studies have identified that characteristics of the donor and recipient characteristics influence short and long term outcomes after transplantation. In this project, we are investigating machine learning methods to develop and validate a kidney donor and recipient risk prediction tool for death, graft loss and the combination of death and graft loss after kidney transplantation, to achieve optimal organ allocation.
2. Phenotyping Donor and Recipients: Current tools using donor characteristics abstracted from administrative data may not adequately capture post-transplant risk for older kidney transplant recipients. We are investigating machine learning based clustering methods to generate unique kidney donor and recipient phenotypes, for different donor and recipient groups, to identify novel combinations of established donor predictors of recipient risk and study the association between the donor and recipient characteristics that impact kidney graft failure.
3. Lipid Management Clinical Decision Support: Knowledge-driven clinical decision support involves the computerization of paper-based clinical guidelines to provide evidence-based recommendations at points-of-care. In this project we computerizing clinical guidelines on lipid management for chronic kidney disease to provide decision support for primary care providers. We are developing a semantic web based framework for modeling clinical guidelines as a Task Network Model that can be executed against patient health profiles to generate personalized recommendations. Our guidelines modeling approach encodes an extensible finite state machine executional semantics using the Notation3 Semantic Web language, which provides powerful features for decisional criteria and queries, and offers integration with the HL7 FHIR standard. 4. Explainable Kidney Graft Survival Analysis: Prognostic modelling using machine learning techniques have been used to predict the risk of kidney graft failure after transplantation. Despite the clinically suitable prediction performance of the models, their decision logic cannot be interpreted by physicians, hindering clinical adoption. We are investigating a model-agnostic XAI approach to kidney transplant survival analysis, in terms of explaining which donor and recipient features contribute most to graft loss and importance of clinical features on outcome modulates over time.
Predicting Clinical Outcomes for Patients Admitted to Intensive Care Unit
Collaborators: Osama Loubani, Tobbias Witter, Karthik Tennankore, Jason Quinn, Calvino Cheng
Funding: NSERC Alliance
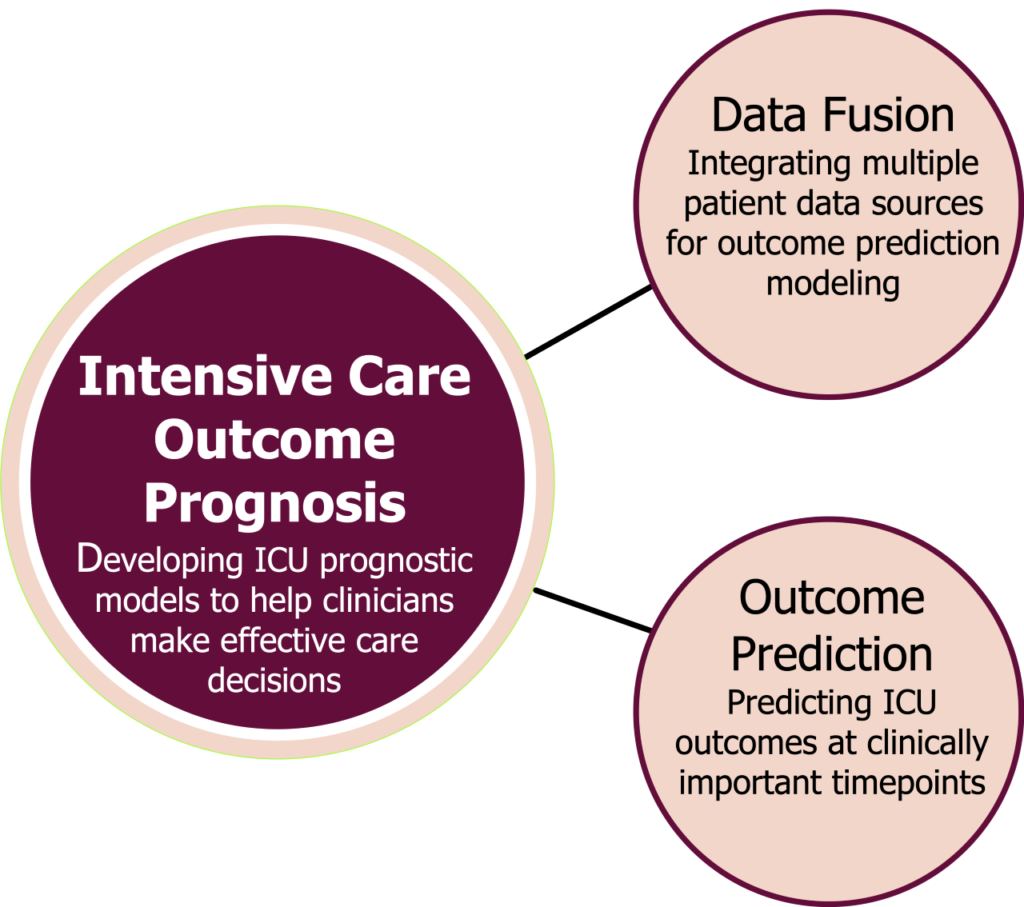
Synopsis: When injury or illness severely compromises vital bodily functions, life-supporting interventions are provided in specialized hospital units called Intensive Care Units (ICUs). The ability to accurately predict the risk of death or clinical deterioration of each patient at various points during their stay in an ICU allows clinicians to offer individualized treatment and intervene more quickly to prevent potentially irreversible injury by allocating resources more effectively. However, currently available prediction scores cannot accurately predict patient outcomes in individual ICU patients at multiple time-points while being treated in an ICU. This project’s objective is to enhance the ICU physician’s decision-making by predicting clinical outcomes across clinically important timepoints during the patient’s ICU stay. We are investigating machine learning models for multimodal data fusion (ICU monitoring, pathology and radiology data), time-series analysis, transfer learning and predictive modeling to develop risk prediction models.
Prediction Models for Arsenic-induced Cancer Risk Using Artificial Intelligence-based Data Analytics Methods for Evidence-based Cancer Prevention and Early Detection
Collaborators: Ellen Sweeney, Jong Sung Kim, Trevor Dummer, Gabriela Ilie
Funding: NFRF, NS Health
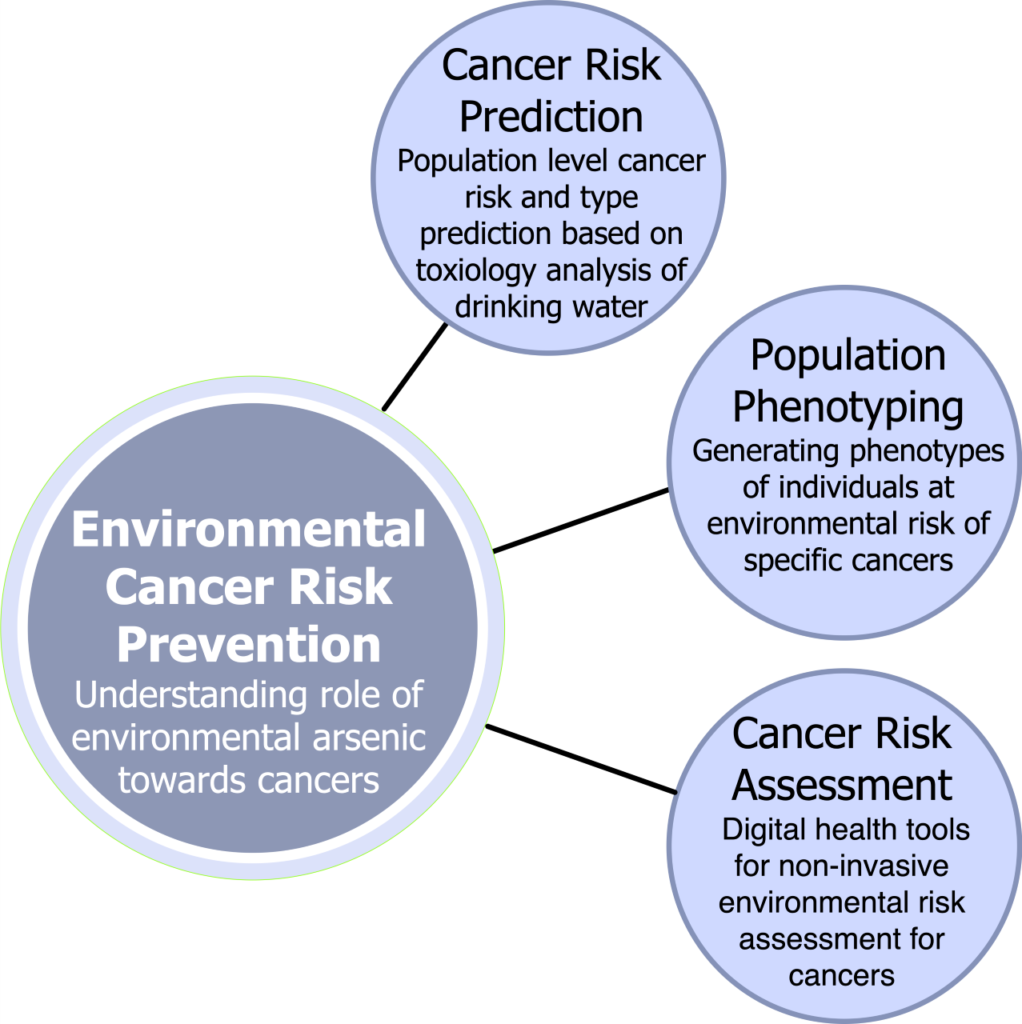
Synopsis: Globally, arsenic (As) contamination of groundwater is widespread. Arsenic has been linked to several types of cancer, including skin, bladder, kidney, prostate, lung, breast and cervical cancer. Based on Canada’s geology, As represents one of the most prevalent environmental carcinogens affecting Canadians. In this research, we seek to reduce the burden of high-fatality cancers caused by As exposure in Canada using the full development spectrum from analytical toxicology, data analytics and population health using toenail biomarkers. We are investigating machine learning methods to (a) profile As species and “metallomes” (entirety of metals) of human toenails implicated in cancer development (b) Develop predictive models for As-induced cancer risk based upon the profiles of As species and metallomes, (c) Assess the relationships between As speciation and cancer using ML methods, and (d) develop prevention strategies for As-induced cancer.
Activity Recognition for Intelligent Assisted Living in Home-based Settings
Collaborators: -
Funding: NSERC
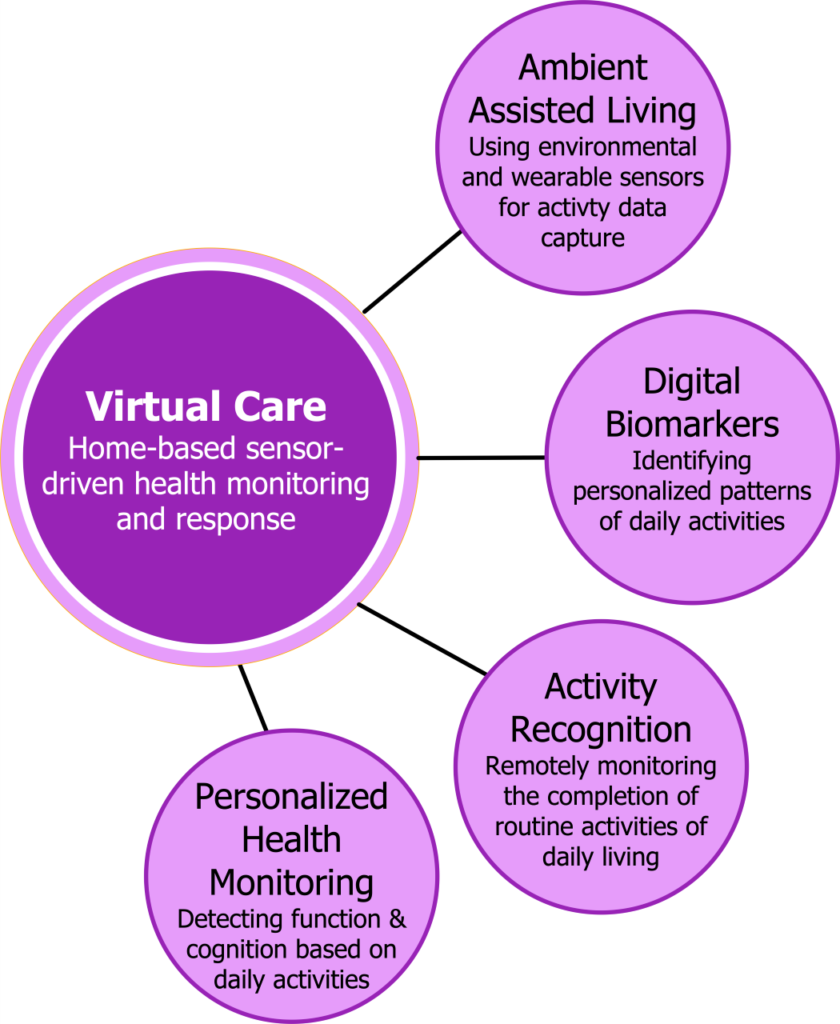
Synopsis: This project aims to develop a novel digital health based intelligent assisted living/virtual care platform that employs ambient intelligence to remotely monitoring an individual’s daily living activities in a home setting to determine health and functional wellbeing. We are investigating AI methods for Activity Recognition (AR) based on remote sensing using wearable and environmental sensors and analyzing the data using A hybrid of knowledgedriven (semantic web) and datadriven (machine learning) AI methods. The AR methods will be used to generate digital biomarkers to assess the ability of an individual to perform dailylife activities. The AR based health assessment will be implemented as digital health based virtual care platform for healthcare providers to remotely monitor their patient’s health status. This project’s intent is to support seniors and patients with dementia, mental health and chronic conditions to safely live in their homes.
AI and Digital Health for Digital Therapeutics Targeting Personalized Healthcare
Collaborators: --
Funding: CIHR, Industry partners
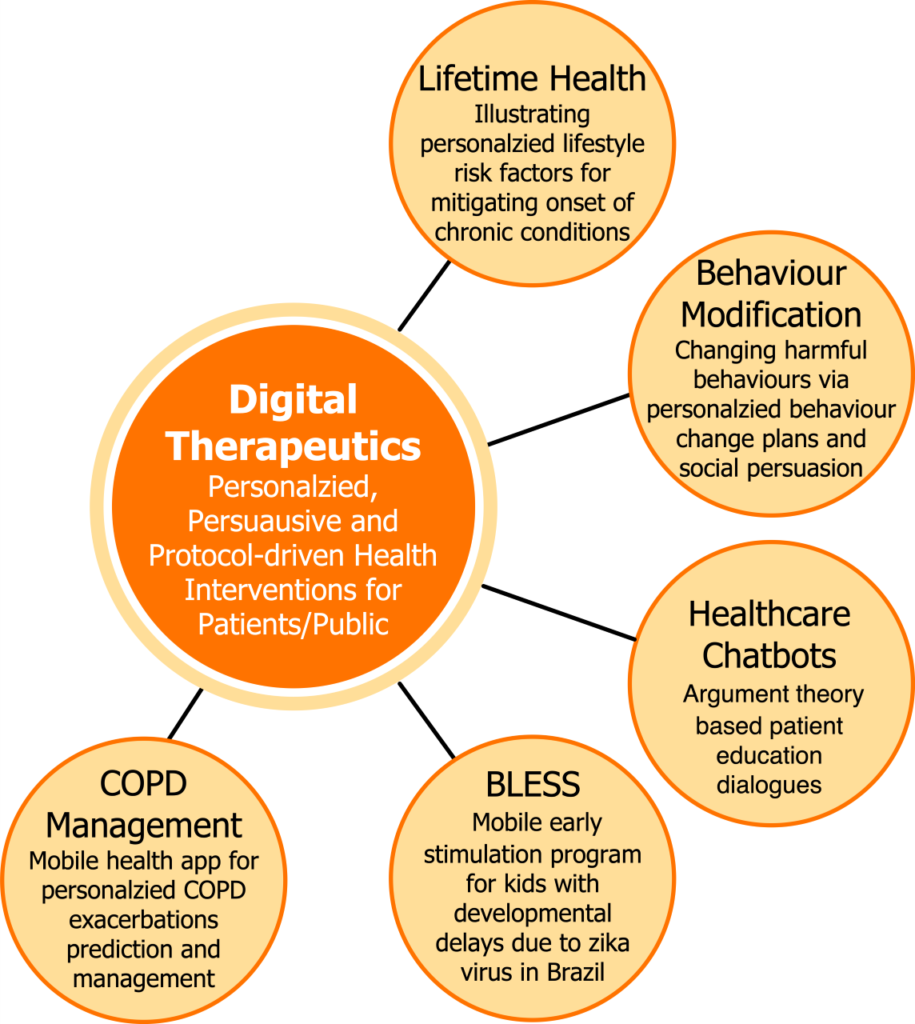
Synopsis: This is a broad research program that focuses on developing innovative digital therapeutics for personalized healthcare targeting patients and the public. The range of projects include:
1. PRISM (Personalized Risk Investigation, Stratification and Mitigation) Platform: Chronic non-communicable diseases, such as cardiovascular disease, diabetes and cancers are regarded as lifestyle-related diseases. Lifetime health is an emerging health paradigm that aims to assist individuals to achieve long-term health targets, and avoid harmful lifecycle choices to mitigate the risk of chronic diseases. The PRISM platform aims to empower Canadian to take personalized measures to prevent the onset of chronic diseases by performing periodic risk assessment, risk monitoring, and lifestyle change to pursue lifetime health. The PRISM platform is designed to provide informative and interactive tools for individuals to (a) assess their cumulative chronic disease risk by providing their health status, and (b) pursue personalized behavioral interventions targeting lifestyle changes to mitigate the noted risk of chronic diseases. At present, we are targeting PRISM to assess the risks for 5 chronic conditions and 8 cancers. The PRISM platform leverages AI, visual analytics and mobile health technologies.
2. Behaviour Modification: Behavior modification interventions help individuals to abandon health-compromising behaviors and adopt health-enhancing behaviors. This behaviour modification project aims to assist chronic disease patients (such as diabetes) to achieve self-management efficacy by helping them overcome barriers to healthy lifestyles. We are developing a personalized behavior modification framework, called Engage, that (a) incorporates a knowledge based ontological model that computerizes behaviour modification theories, such as cognitive behavioural therapy, social cognitive theory and others, (b) employs social modeling techniques to encourage individuals to pursue their behaviour modification plans, and (c) uses logical reasoning over the knowledge model to generate personalized action plans as per the patient’s health and behavioral profile.
3. Virtual Care: This project is in collaboration with researchers in Brazil where the Zika virus outbreak caused microcephaly in newborns, which affects cognitive development delays. Treatment for developmental delays in children between 0-3 years comprises a standardized exercise program—known as the Early Stimulation Program (ESP) that targets cognitive and motor function development. To improve timely access to ESP treatment, we are implementing a home-based ESP delivery approach that remotely trains the parents of affected children about how to perform the ESP at home. In this project, we are developing, implementing and evaluating a digital health based ESP therapy intervention called the BraziLian Early Stimulation System (BLESS) that incorporates (a) mobile health app that allows parents to receive the prescribed ESP videos to help them administer ESP in a home-based setting, and (b) an intelligent ESP therapy planning and decision support tool to support therapists to design an evidence-based ESP based on the child’s needs. 4. Patient Educational Chatbots: The successful long-term treatment of chronic conditions relies on the empowerment of patients and caregivers to self-manage non-acute symptoms. This requires comprehensive knowledge on the illness, hence, chronic disease patients are typically provided with educational resources. We are pursuing a novel approach for patient education that involves dialogue by design, where credible dialogues are dynamically formulated directly from the structured arguments of the dialogue, thus removing the need for predefined linear discourses, as is the case in most health-related dialogue systems. For dialogue construction, we have computerized the Toulmin model of argument by applying semantics-based knowledge engineering to engineer an argument ontology to semantically encode patient education material.
Digital Health and Artificial Intelligence based Platform for Early Chronic Disease Risk Prediction to Improve Population Health
Collaborator: Ellen Sweeney, Sean Christie, Osama Loubani, Karthik TennankoreFunding: CIHR
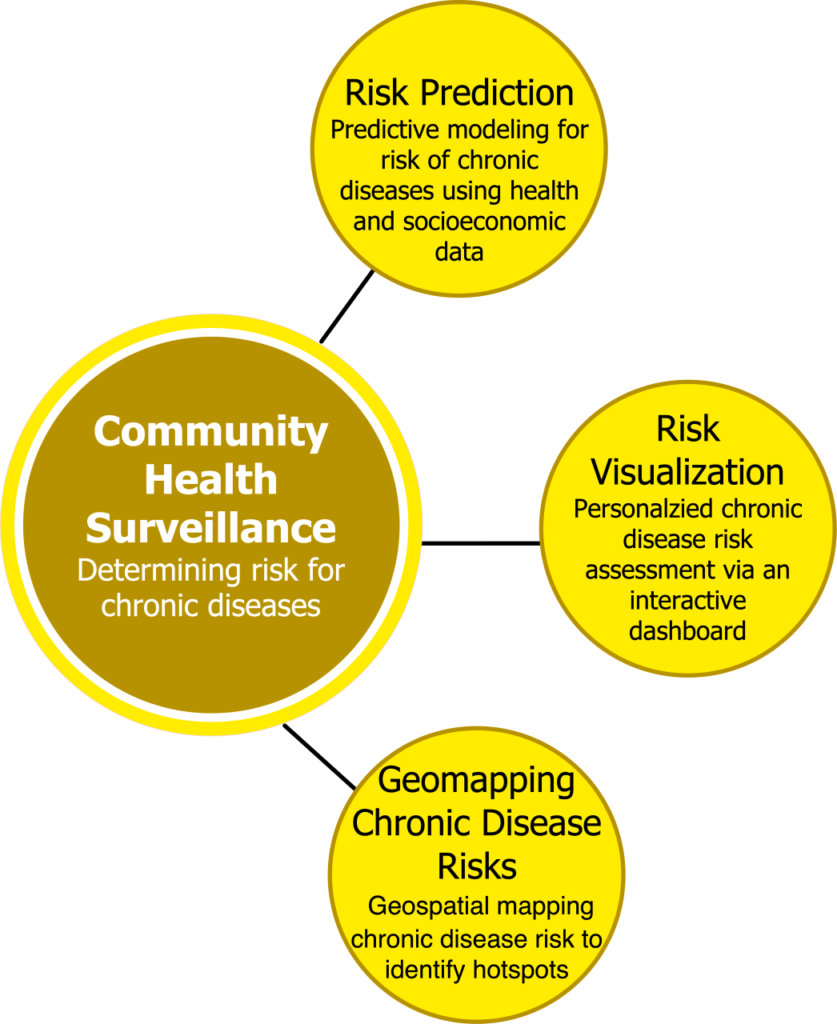
Synopsis: Lifecourse health is an emerging paradigm that aims to assist individuals to achieve long-term health targets, and avoid detrimental choices that may affect their long-term health. The objective of this population health project is to develop and evaluate an innovative Digital Health and Artificial Intelligence (AI) enabled intervention to assist citizens in preventing the onset of behaviour-related chronic diseases.
We are developing an online public health platform—i.e., Personalized Risk Investigation, Stratification and Mitigation (PRISM)—to engage and educate citizens to perform chronic disease risk self-assessment, monitoring and predictions to prevent the onset of chronic diseases. We will use AI methods to (a) computerize multiple chronic disease risk assessment tools to provide multimorbid chronic disease risk assessments; (b) predict the future risk for chronic diseases by analyzing population health data from Atlantic Partnership for Tomorrow’s Health (PATH); and (c) illustrate to individuals their health status and chronic disease risk assessments using a digital health based online personalized health monitoring dashboard that employs visual analytics methods to visualize behavioral risk factors to multimorbid risks.
Health Knowledge Management
Collaborators: --
Funding: Industry partners
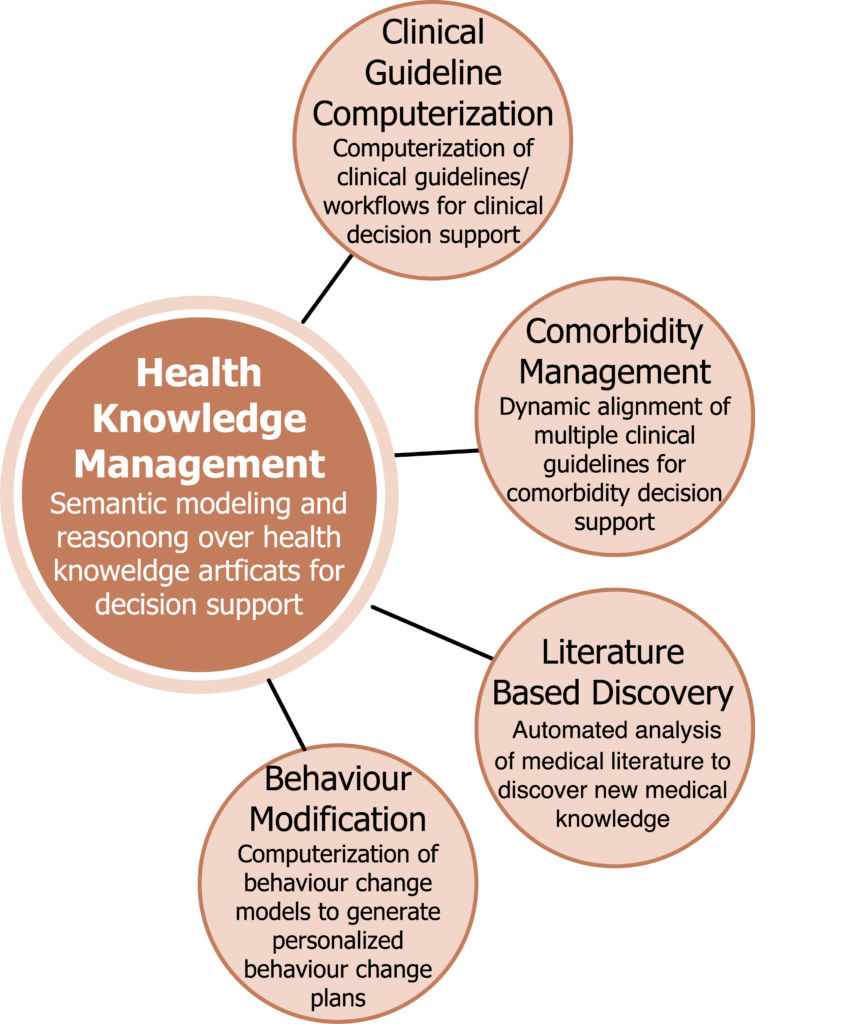
Synopsis: This research program focuses on investigating knowledge-based methods, particularly semantic web, ontologies, knowledge graphs, for health knowledge modeling to develop innovative clinical decision support systems and literature based discovery systems. The range of projects include:
1. Clinical Guideline Computerization: Clinical Practice Guidelines (CPG) contain evidence-based recommendations to guide clinicians in diagnosis, prognosis, and treatment of a medical condition. We are developing knowledge modeling solutions, using ontologies, to computerize/digitize health knowledge (such as such as clinical guidelines, psychosocial theories, clinical workflows, etc.) as knowledge models that can be reasoned with patient data to deduce context-sensitive, personalized, evidence-based decision support recommendations.
2. Comorbidity Management: Clinical management of comorbid conditions is highly complex, as it demands administration of therapeutic interventions for multiple conditions whilst avoiding adverse interactions (e.g., drug-drug or drug-disease), duplicate investigations. Handling comorbidities using computerized clinical guidelines is a challenging task as it demands the integration of multiple disease specific clinical guidelines. We are developing a knowledge modeling approach, using semantic web based methods, to dynamically integrate two or more comorbid clinical guidelines at execution-time to effectuate a clinically safe and efficient comorbid clinical workflow. Our approach handles the dynamically evolving condition of the patient, and as such we care investigating temporal logic to handle temporal aspects of clinical tasks (e.g., sequential relations, task durations), transaction logic and guideline integration ontology to facilitate the integration of multiple comorbid guidelines
3. Literature Based Discovery: Knowledge Synthesis to Discover Novel Mechanistic Associations From Medical Literature
Collaborators: —
Synopsis: The published biomedical literature contains a significant volume of implicit knowledge that if discovered can contribute to incidental finding about complex issues such as gene-disease associations, cancer pathogenesis, and new indications for existing drugs (i.e., drug repurposing). We are developing a medical literature based knowledge discovery framework that integrates knowledge graphs, semantics driven text analysis, systems medicine and reasoning methods to synthesise, analyze and discover new knowledge from medical literature. Our focus is to discover new knowledge associating (i) cancer risk factors with incidence of cancers, (ii) COVID-19 with chronic conditions in terms of long covid, and (iii) cancer drug repurposing.